How AI-Driven Protein Folding Is Rewriting the Source Code of Life
Decoding Creation: How AI Protein Folding & Alpha Fold are accelerating Drug Discovery
Every protein in your body — every enzyme digesting your breakfast, every antibody hunting a virus, every structural fibre holding your tissues together — is, at its core, a long chain of amino acids. There are only twenty of these building blocks, but the chain can be hundreds or thousands of units long, and each chain folds into a unique, intricate three-dimensional shape. That shape is the protein’s function
Think of a straight piece of wire. By itself, it can’t do much. But if you bend it into the shape of a paperclip, it suddenly has a very specific function: holding papers together. If you bend it into a hook, it can catch a fish.
The shape you give the wire completely determines its job. Proteins inside your body work in the exact same way.
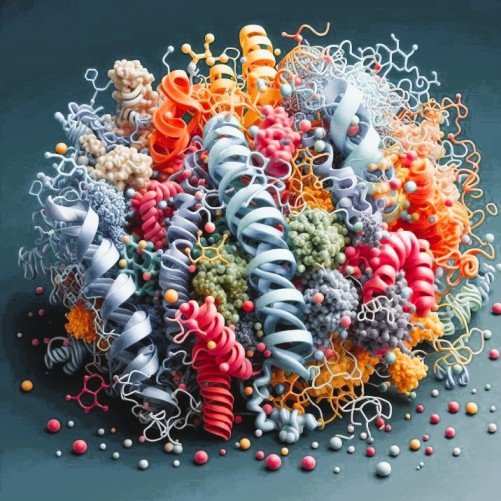
- The Building Blocks (The String of Beads)
Protein doesn’t have a shape at the start. Amino acids, which are tiny building blocks, make up long, straight chains that start their lives. - Protein folding (molecular origami)
You can’t do much with a flat piece of paper until you fold it into an origami crane or a paper airplane. The same goes for amino acids; it has to be folded to work.
The “beads” have different chemical properties that make the chain naturally twist, scrunch, and fold itself into a very complicated three-dimensional shape. Some like water, some hate it, some are positively charged, and some are negatively charged. This exact way of scrunching is called protein folding. - Structure = Function (The Lock and Key)
- The relationship between folding and function boils down to one golden rule in biology: Structure dictates function. The exact 3D shape the protein folds into determines exactly what it can do in the body.
One important rule in biology is that structure determines function. This is what folding and function have in common. The precise three-dimensional configuration of the protein dictates its functional capabilities within the organism.
Most proteins do their jobs by attaching to other molecules. The protein’s folded shape has to fit perfectly with the shape of its target, like a key and a lock.
- The “Scissors” Shape: If the protein folds into a shape with a small, sharp pocket, it could work like an enzyme, which is a pair of molecular scissors that grabs your food and cuts it up so you can digest it.
- The “Tube” Shape: If it folds into a hollow cylinder, it might get stuck in a cell wall and do something.
- If it folds into a hollow cylinder: It might embed itself into a dell wall and act as a secure tunnel, allowing specific nutrients to pass in and out.
- The “Rope” Shape: If multiple proteins fold into long, stiff, intertwining cables, they become structural proteins, like the keratin that gives your hair and nails their incredible strength
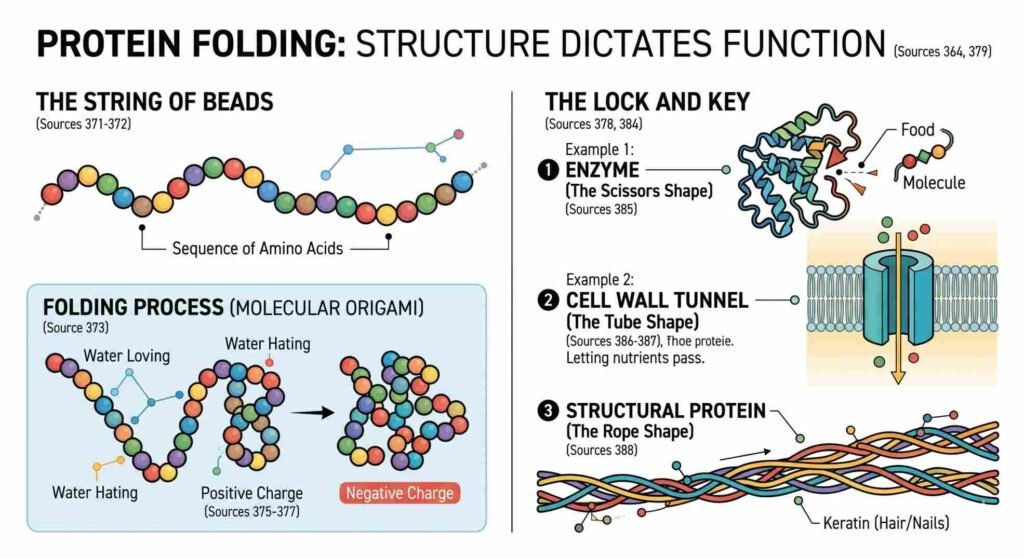
The 50-Year Puzzle That a Machine Solved in Minutes
Scientists worked on what is now called the “protein folding problem” for more than fifty years. The question was, if you only have a raw sequence of amino acids, can you guess what the protein’s three-dimensional structure will be? X-ray crystallography, nuclear magnetic resonance spectroscopy, and cryo-electron microscopy are all very powerful but very slow. It could take months or years of hard work in the lab and cost hundreds of thousands of dollars to figure out the structure of just one protein. The global scientific community had put about 100,000 experimentally solved structures in the Protein Data Bank by 2020, after decades of work.That sounds great, but it’s only a tiny part of the billions of protein sequences that scientists have found in all living things on Earth.
Then, in just a few amazing years, AI changed everything
Alpha Fold: The AI That Stunned the World
In December 2020, DeepMind’s AlphaFold 2 entered the 14th Critical Assessment of Structure Prediction (CASP14) competition — the field’s biennial Olympics — and demolished the competition. AlphaFold 2 could predict a protein’s structure from its amino acid sequence alone, in GPU minutes rather than laboratory months, with accuracy rivalling experimental measurements. The programme achieved amazingly high accuracy scores.
The numbers are staggering. By July 2022, the AlphaFold team had predicted the structures of virtually all 200 million proteins then known to science and made them freely available through the AlphaFold Protein Structure Database. By late 2025, the AlphaFold 2 paper in Nature had been cited nearly 43,000 times, and over two million researchers from 190 countries had used the database. A process that once took a doctoral student’s entire career could now be done before lunch.
AI-Driven Protein Folding Is Transforming Modern Biology
AlphaFold 3: From Proteins to the Molecular Universe
Released in May 2024 by Google DeepMind and Isomorphic Labs, AlphaFold 3 took a dramatic leap beyond its predecessor. While AlphaFold 2 focused on individual protein chains, AlphaFold 3 can predict the joint three-dimensional structure of entire biomolecular complexes — proteins interacting with DNA, RNA, small-molecule drugs, ions, and post-translational modifications, all in a single computation.
The effect on the ground is huge. In 76% of cases, AlphaFold 3’s predicted binding poses for protein-ligand interactions are within two angstroms of experimental structures. This is about twice as accurate as the best previous specialized docking tools. This changes virtual drug screening: drug companies like Eli Lilly and Novartis can now use computers to screen thousands of candidate molecules against target proteins, finding hundreds of promising drug candidates before they even touch a test tube. It used to take six months of crystallography, but now it can be done in forty-eight hours of computing.
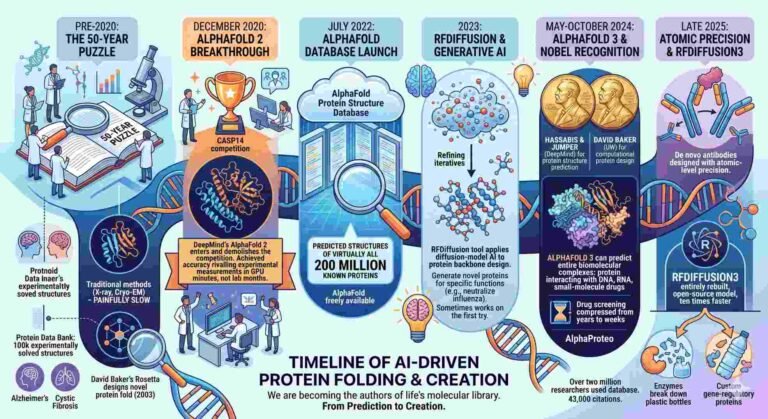
AlphaFold 3 uses a completely new architecture based on diffusion models, which are the same type of AI that powers tools that make images. It starts with a cloud of atoms and gradually refines their positions into a coherent three-dimensional structure.
The model is not merely faster; it represents a conceptually different approach to understanding molecular biology, treating the cell’s molecular machinery as an integrated system rather than a collection of isolated parts.
From Prediction to Creation: The Generative AI Revolution in Biology
If AlphaFold reads biology’s existing blueprints, the next generation of AI tools is writing new ones. This is the frontier of de novo protein design — creating proteins that have never existed in nature, tailored to perform specific functions on demand.
David Baker’s laboratory at the University of Washington has been the epicentre of this revolution. In 2003, Baker’s Rosetta programme designed a completely novel protein fold from scratch — a feat many believed impossible. Since then, the lab has produced an astonishing menagerie of designer proteins:
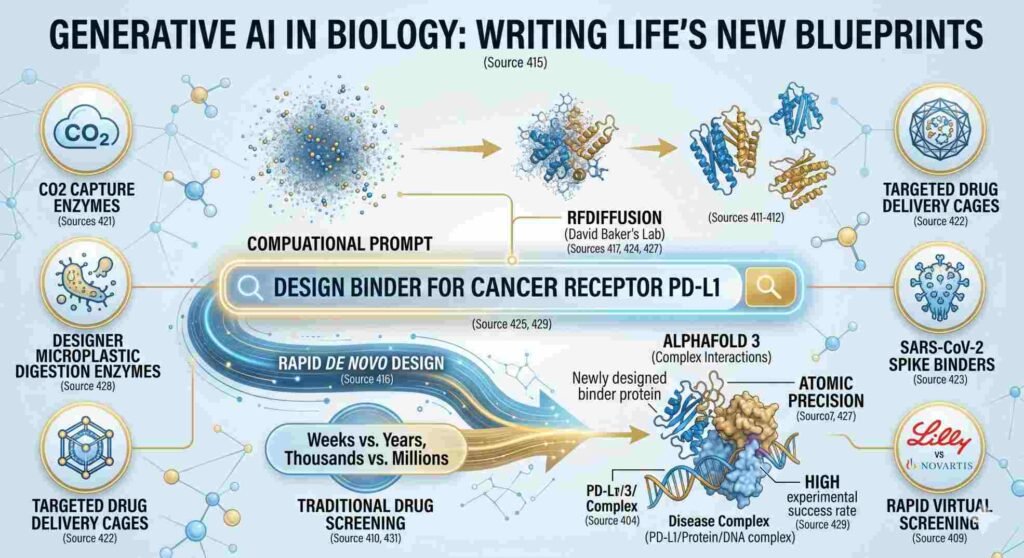
- peptides that neutralise multiple influenza strains,
- enzymes that help microbes capture atmospheric carbon dioxide,
- self-assembling molecular cages for targeted drug delivery, and
- binders that latch onto the SARS-CoV-2 spike protein to block infection.
The pace of innovation accelerated dramatically with RFDiffusion This tool applies diffusion-model AI — the technology behind image generators like DALL-E — to protein backbone design. A scientist specifies a desired function (“bind to this cancer receptor,” “scaffold this enzyme active site”), and RFDiffusion generates entirely novel protein structures optimised for that purpose. A dramatic improvement over older approaches that required screening tens of thousands of candidate molecules.
In December 2025, the team released RFDiffusion3, an entirely rebuilt, open-source model that operates at the level of individual atoms and is ten times faster than its predecessor. It can design proteins that interact with DNA, small molecules, and other non-protein targets — opening the door to custom gene-regulatory proteins, designer enzymes for breaking down microplastics, and biosensors for environmental monitoring.
Meanwhile, Google DeepMind’s AlphaProteo, introduced in 2024, takes a complementary approach: given a target protein and a desired binding site, it generates novel binder proteins with high experimental success rates across diverse targets including cancer-related proteins like VEGF-A and PD-L1. In head-to-head comparisons, AlphaProteo matched or exceeded RFDiffusion’s success rates on multiple targets, demonstrating that the AI protein design field is now a rapidly advancing competitive ecosystem.
From Sequence to Structure: How AI Is Solving Biology’s Biggest Puzzle..it is going to lead drug discovery at the speed of thought… The speed of AI design offers a tantalising prospect: With AI tools, they can predict the structure of the virus’s key surface proteins, design custom binder molecules that block infection, and begin laboratory testing — all within weeks rather than the months or years that traditional approaches demand. Baker’s lab demonstrated this possibility. designer proteins could improve carbon capture by engineering enzymes that more efficiently convert atmospheric CO₂ into stable compounds.
With the fast pace discovery of AI Protein folding and structure prediction Are We Becoming Architects of Life? Given these godlike powers over the molecular building blocks of life, a provocative question naturally arises — one that has fascinated humanity since the myth of Icarus, the sketches of Leonardo da Vinci, and the wild imaginings of science fiction: could we grow wings on a human being?



